Dynamic response of liver to varying stiffness levels
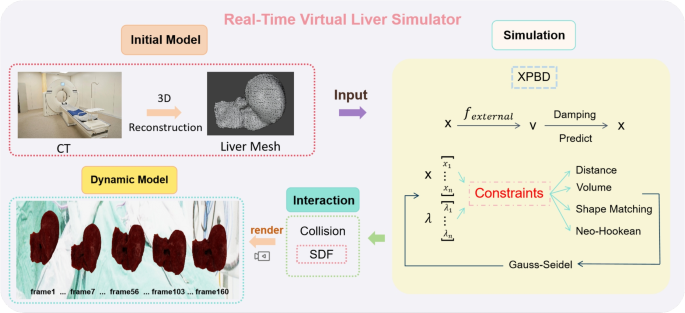
Deformations are common in simulations of medical procedures, such as organ manipulation, surgical transection, and laparoscopic interventions. In this study, we used real-world patient CT scan data from the publicly available 3D-IRCADb-01. The liver data were three-dimensionally reconstructed using 3D Slicer software, and a volumetric tetrahedral mesh was generated using Delaunay triangulation in the CGAL library. All experiments were conducted using an NVIDIA GeForce RTX 4090 GPU. Our simulator dynamically adjusts the alpha of various constraints to model livers with different stiffness levels, accounting for variations due to age, population, and diseases like cirrhosis or fatty liver, which significantly alter liver stiffness. As liver stiffness is a critical biomechanical marker of disease progression, this visualization is invaluable for understanding and simulating disease impact.
However, conventional ultrasound presents several drawbacks in the preliminary diagnosis of liver fibrosis, including low sensitivity, subjectivity, and limitations in both qualitative and quantitative assessments. To address these issues, we used shear wave elastography (SWE), a non-invasive and effective method for quantitative data collection. SWE generates transverse shear waves via ultrasound excitation within the tissue, detected by ultra-fast imaging technology. The tissue’s elasticity is visualized in real time using color-coded maps, enabling the quantitative measurement of Young’s modulus and Poisson’s ratio. These parameters provide a direct, intuitive measure of tissue stiffness, facilitating the assessment of liver hardness and fibrosis extent. The formulas for Young’s modulus Ε and Poisson’s ratio ν are presented in Eqs. (15)–(16) by Lamé parameters, \(\lambda\) and \(\mu\):
$$\begin{aligned} & \lambda = \frac{E\nu }{(1 + \nu )(1 – 2\nu )}, \end{aligned}$$
(15)
$$\begin{aligned} & \mu = \frac{E}{2(1 + \nu )}. \end{aligned}$$
(16)
Young’s modulus quantifies the rigidity of a material, with higher values indicating greater resistance to deformation. An increase in Young’s modulus and decrease in Poisson’s ratio reflects the progressive stiffening of the liver, which correlates with the advancement of fibrosis. Quantitative measurements of liver stiffness across different individuals are typically expressed in kPa. Table 1 provides four data quantiles selected from publicly available detection results of diverse health populations. For clarity, the results are organized into four rows, each corresponding to a distinct stiffness configuration using Young’s modulus and Poisson’s ratio, we calculated compliance denoted as \(\alpha\) to dynamically adjust four constraints in the physics engine.
Figure 2 illustrates the deformation process of the liver model under different constraint conditions. As part of our real-time virtual liver simulator, we captured frames during the experiment to display this process. Each column in the figure represents screenshots taken at different stages of the simulation. Each row in the figure represents a specific set of stiffness settings, ranging from softer (top row) to harder livers (bottom row). By gradually increasing the stiffness in the simulations, we observed variations in liver deformation under identical external forces. These external forces include gravity acting downward, friction with the ground, and elastic forces resulting from collisions. We defined the deformation task as the process in which the liver begins to deform from the moment it is subjected to external forces and collides with the ground, continuing until it returns to its original form without undergoing further significant deformation. Our experimental setup was designed to capture the dynamic changes in the liver when subjected to external forces, simulating physical interactions that could occur in real-life scenarios. Through detailed visual analysis, our study not only reveals the realism of the liver physical model but also provides valuable data supporting future biomechanically-based medical diagnosis and treatment planning.
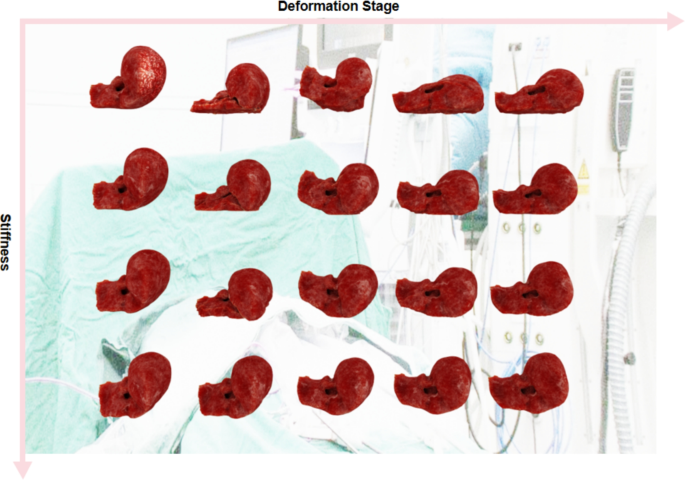
Deformation of the liver model under varying constraint conditions, as demonstrated in selected frames from our simulator. Each row systematically showcases a distinct stiffness configuration, ranging from more compliant (top row) to increasingly rigid (bottom row), highlighting the impact of these conditions on liver deformation.
Our engine not only integrates the physical constraints of the Neo-Hookean model but also allows for the substitution of other material models to customize the elasticity controls. Furthermore, the integration of Shape Matching constraints and the tunability of the Distance and Volume constraints enhance our model’s flexibility, extensibility, and fidelity, offering significant improvements over traditional PBD and FEM approaches. Our study also demonstrates the capability of constructing a liver simulator for medical visualization based on real patient data. The simulator achieves highly realistic physical and visual effects, aligning with the truth of computer graphics: if it looks right, it is right36.
Comparative experiment
In our study, we specifically focused on comparing the real-time performance and visual realism of different simulation methods. We evaluated three distinct methods: traditional PBD, our liver simulator, and FEM on an identical liver mesh. The results shown in Table 2 record the average simulation times for handling the aforementioned defined deformation tasks and FPS during the runtime of the liver simulator. Notably, although our simulator recorded a slightly slower time than PBD, it significantly outperforms FEM in both runtime and FPS, clearly demonstrating its superior real-time performance capabilities. This marginal increase in processing time is justified by the significant enhancements in simulation complexity and physical realism offered by our simulator, establishing it as an effective solution for real-time medical simulation applications.
Visual Comparison of Liver Simulators: Sequential images from left to right show the simulation results generated by PBD (Yellow), Our Simulator (Red), and FEM (Blue).
Striking Visual Comparison of Liver Simulators: Our simulator (outlined in orange) is overlaid with PBD (first image) and FEM (second image), clearly illustrating the differences in deformation characteristics at the point of maximum deformation on an identical liver mesh.
XPBD incorporates advanced handling of physical properties, such as accurately simulating elasticity and dissipative energies. By introducing Lagrange multipliers and well-defined constraint functions, it ensures the fulfillment of positional constraints while precisely reflecting the physical responses of biological tissues like the liver, including elastic deformations and compressive characteristics. Figure 3 shows a visual comparison of liver simulators and in Fig. 4, our results, outlined in orange, are overlaid with PBD (first image) and FEM (second image) for clearer comparison. This overlay distinctly reveals that PBD produces unrealistic mesh protrusions and deformations caused by violent ground collisions, while FEM shows similar but slightly smaller deformations compared to our results, emphasizing the robustness and visual realism of our simulator. Unlike PBD, which lacks physical realism and does not incorporate physical properties, especially in handling complex biomechanical attributes such as the liver’s nonlinear elasticity and viscoelasticity, our simulator addresses these critical deficiencies. This added complexity is essential for achieving accurate results in medical simulations that demand high physical fidelity.
Severe mesh distortion in soft-body simulations frequently leads to non-convergent solutions, fundamentally revealing inherent limitations in numerical stability and precision when conventional simulation algorithms process large deformations. Although mitigation strategies such as Laplacian smoothing have been proposed, they incur critical trade-offs: iterative mesh smoothing inherently increases computational overhead and time consumption, with excessive iterations potentially introducing artificial model contraction. More importantly, such geometry-focused corrections neglect underlying physical motion characteristics, manifesting as numerical instabilities rooted in algorithmic precision limitations. To quantitatively evaluate mesh integrity, we define a smoothness metric S as:
$$\begin{aligned} S = \frac{1}{n} \sum _{i=1}^{n} \frac{V_{\min ,i}}{V_{\max ,i}} \end{aligned}$$
(17)
where n is the number of adjacent element pairs, and \(V_{\min ,i}, V_{\max ,i}\) represent the smaller and larger volumes of adjacent tetrahedral elements, respectively. This dimensionless metric ranges from 0 to 1, with higher values indicating better volumetric consistency. Our simulator addresses these limitations through the integration of volumetric constraints and physical material model constraints. Conventional finite element methods often encounter similar instability challenges when they employ inappropriate constitutive models, particularly in complex biomechanical scenarios. Experimental validation confirms our methodology’s enhanced robustness over conventional approaches, establishing a reliable computational foundation for dynamic mesh models simulating anatomical deformations. Table 3 presents smoothness values across the three methods during maximum deformation. Our simulator demonstrates superior mesh integrity compared to both PBD and FEM approaches, with higher mean smoothness and lower standard deviation, indicating more consistent element volumes throughout the deformation process. This quantitative advantage directly correlates with the visual differences observed in Fig. 4, where PBD exhibits problematic mesh distortions during collision events.
In contrast, while FEM offers slightly higher visual realism than PBD, its computational efficiency is lower, making it unsuitable for applications requiring real-time feedback. Our simulator achieves visual realism comparable to FEM while maintaining computational efficiency similar to PBD, effectively bridging the gap between these approaches. Therefore, our simulator is well-suited for use in virtual surgery and other scenarios that require both high-fidelity visualization and real-time feedback.Despite the advantages of our XPBD-based approach, it is important to acknowledge certain limitations specific to liver simulation applications. While XPBD effectively addresses the iteration count and time step dependency issues of traditional PBD33, our implementation still faces challenges in accurately modeling extreme anisotropic behavior that may be present in diseased liver tissue with significant fibrosis. Additionally, the accurate simulation of discontinuities that occur during surgical cutting procedures requires special handling beyond standard XPBD constraints, potentially necessitating adaptive remeshing techniques that could temporarily impact performance37. For scenarios involving complex fluid-structure interactions, such as blood flow within the liver’s vascular network, our current constraint system would require extension with specialized fluid handling capabilities. These limitations do not significantly impact the primary applications of preoperative planning and training, but represent important considerations for future development toward comprehensive surgical simulation environments. The extensible nature of our constraint-based framework provides a foundation for addressing these challenges while maintaining real-time performance.
Applications and future prospects
The real-time virtual liver simulator significantly improves the accuracy of preoperative planning38 and intraoperative navigation39, especially in ultrasound-guided biopsies40, substantially improving surgeons’ ability to precisely locate complex liver structures, such as blood vessels and bile ducts. This optimization of surgical strategies reduces risks and increases success rates. Additionally, this technology provides a safe simulation environment in the field of medical education and professional training, allowing medical students and interns to improve their clinical operation skills through repeated practice. It also supports high-precision training for robot-assisted surgeries, helping surgeons master the skills needed to operate sophisticated robotic devices. Furthermore, liver simulation technology demonstrates its broad potential in applications such as liver transplant assessment and planning, as well as postoperative rehabilitation training and evaluation, all of which rely on accurate and real-time simulation capabilities. Particularly in telemedicine, this technology supports remote guidance and decision-making for complex surgeries by physicians in remote areas, enhancing surgical success rates and patient survival, especially valuable in resource-limited settings.
Building on this array of applications, our proposed methodology showcases substantial potential in liver simulations, highlighting the remarkable scalability and robustness of our virtual liver simulator. This scalability positions our work within the emerging paradigm of human body digital twins, which aims to create comprehensive digital representations of human physiology for personalized healthcare. A recent roadmap for developing human body digital twins describes a progression from isolated cross-sectional models to fully integrated physiological systems41. While pioneering efforts have focused on individual organs, such as GPU-accelerated digital twins of the human heart42 and advanced 3D reconstruction of tissues43, or specific physiological systems such as wearables-based prediction of motion intention through the nervous system44, the future lies in integrating these specialized components into comprehensive frameworks. Our liver simulator contributes a critical element to this larger vision, where multiple organ simulations could eventually be combined to model their interdependent relationships. As human body digital twin technology advances, our high-fidelity liver simulation methodology offers a valuable building block that could help bridge the gap between organ-specific models and fully integrated physiological systems. This approach is not only effective for liver simulations but is also adaptable for simulating other organs, such as the lungs11, heart45, and colon46. We are confident that this method is suitable for all organ simulations and broader medical scenarios, presenting a promising direction for future development. However, it also presents certain limitations; due to the unique biomechanical properties of different human organs, it necessitates the flexible selection of appropriate constraints to ensure that the simulations more accurately replicate the physical realities of the real world. In reality, there are even more complex scenarios. For instance, we have not yet considered situations where the liver may have tumors, causing significant changes in its properties and shape. This would complicate the boundary conditions, making them harder to manage. Extremely complex liver geometries and internal structures could also lead to potential inaccuracies in the simulator. The use of more specific and numerous constraints is among the areas where future improvements could be made.
Future research directions will further expand the application scope and enhance the precision and speed of liver simulation technologies. On one hand, surgical simulators integrated with AR have emerged as cutting-edge tools for surgical training and planning. This technology overlays virtual data onto actual operative environments, providing seamless visual and physical feedback to surgeons, thus enhancing surgical precision and efficiency, and allowing for the optimization of surgical strategies under conditions closely resembling actual scenarios. On the other hand, employing deep learning to optimize the solution processes of physical equations represents a critical direction for future development. By accelerating the computation of complex physical equations using deep learning algorithms, both simulation speed and accuracy are improved, reducing the reliance on high-performance hardware and facilitating the broader deployment of liver simulation technology across diverse medical environments. These research initiatives are anticipated to foster significant advancements in surgical planning, medical education, and telemedicine, thereby enhancing the quality and efficiency of global healthcare services.
link